DNA Fingerprinting
Since childhood, we have been enjoying the thrilling crime dramas where often cases are closed with DNA tests confirming the suspect as guilty. Have you ever wondered what goes on in those testing labs which play such a crucial role in solving crimes?

The procedure used by the forensics team to identify the criminals from their DNA is called DNA Fingerprinting or DNA profiling. Any two people in the world have on an average 99.9% same DNA. The remaining 0.1% is what makes us unique (unless you are an identical twin). Although this might sound like a small amount, this means that there are around 3 million base pairs that are different between two people. So this process analyses the remaining 0.1%. It is a sensitive procedure and only requires a few skin cells or hair root or few drops of blood or saliva because almost all of our cells contain our DNA.
History Behind DNA Fingerprinting
Alec Jeffreys and his team at Leicester University discovered this method on September 10th , 1984. They had been working for seven years to see if it would be possible to differentiate people using their DNA. They had been looking at regions of human genome called VNTRs which stands for Variable Number Tandem Repeats. These can be thought of as a repeating string of bases. The strings consist of 10-80 bases and the number of repetitions can be as high as 30.
These sequences are inherited along with the rest of the DNA from parents. VNTRs occur within the parts of the genome called non-coding regions, which are not used to encode proteins. Therefore genetic changes called mutations and duplications (which are copies of the DNA sequence) take place much more often within the VNTRs than in other parts of the genome. This means that a person is likely to have a similar number of repeats to their close relatives than people more distantly related to him/her and complete strangers will have a totally different number and therefore different length VNTRs. It is these differences that can be exploited to tell one person from another gene ethically.
In 1984, Jeffreys was looking at the results of an experiment he carried out to see if a probe he created to pick up one particular VNTR sequence could show variations in the length of the sequence repeats in DNA of could show variations in the length of the sequence repeats in DNA of his lab assistant and parents.

A clear pattern was visible confirming that he had inherited repeats of different lengths from each parent and thus the world of DNA fingerprinting was born.
Initially, Jeffreys lab was the only place in the world which offered DNA fingerprinting services. Later in 1987, the process was commercialized.
Modern DNA Fingerprinting
Nowadays rather than VNTRs, sequences called STRs (Short Tandem Repeats) are used. These are similar to VNTRs but smaller and stand better to degradation of DNA over time which makes them much more useful for investigating crimes. STRs are sections (or loci) of chromosomes where instead of gene consisting of a long sequence of bases, there are much shorter sequences of three or four or five bases.

At the places on the chromosomes es on the chromosomes where we find these STRs, there are areas that vary in a number of repeats. areas that vary in a number of repeats. DNA profiling only looks at these STRs. A cell sample is collected which could be from some blood at a crime or a swab from the inside of someone’s cheek for example. The DNA is then extracted from the sample. Many copies of this DNA may be made using the Polymerase Chain Reaction (PCR). PCR copies and amplifies specific target DNA sequences, in this case, STRs. Special enzymes called restriction endonucleases are used to cut the DNA up into different sized pieces. The DNA samples are then put into wells in a special gel called agarose for the process of gel electrophoresis which separates the DNA fragments by size.
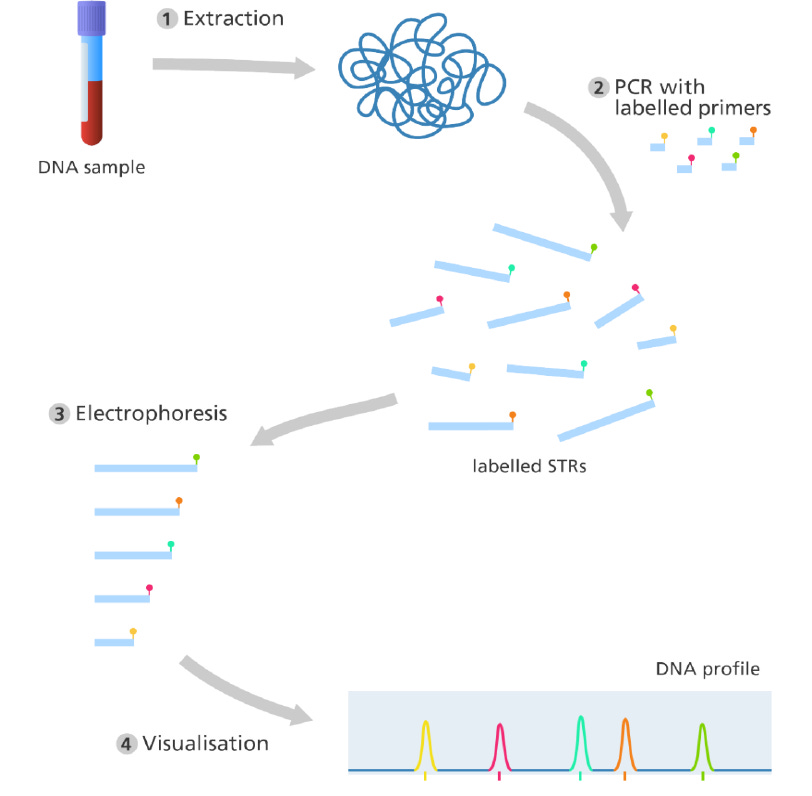
DNA is negatively charged, so it will move through the gel towards the positive electrode. The larger, longer STR repeats which are made up of long strings of DNA letters are heavier so they move slowly through the gel than shorter lighter repeats. So by running the electric current through DNA for a certain period of time, the different sized pieces of DNA will spread out through the gel. They can be revealed using a dye to label them up producing a line called a band on the gel. The pattern is then transferred to a nylon sheet southern blotting. The pattern of these bands showing the presence of different sized repeats is the genetic fingerprint of a person.

Only one person in every 10 million (10,000,000,000,000) will have a particular STR profile. Therefore it is extremely unlikely you will share the same profile as someone else unless you are an identical twin. The UK was the first country to set up a national database of DNA profiles in 1995.
DNA profiling is also used in paternity tests. We inherit half of our DNA from each of our parents. Which is also the case for STRs (or VNTRs as shown in the second figure). We inherit repeats of different lengths from each parent which can be identified by DNA profiling.
An Article By : Danush Pavan Teja
DISCLAIMER
Shaastra TechShots’ publications contain information, opinions and data that Shaastra TechShots considers to be accurate based on the date of their creation and verified sources available at that time. It does not constitute either a personalized opinion or a general opinion of Shaastra or IIT Madras. The information provided comes from the best sources, however, Shaastra TechShots cannot be held responsible for any errors or omissions that may emerge.



