DNA/Molecular Computing

What is DNA Computing?
Molecular computing is a branch of computing that uses DNA, biochemistry, and molecular biology hardware, instead of traditional silicon-based computer technologies. Instead of running software on a traditional computer, some scientists are now trying to replace the silicon chip with test tubes, liquids, and even living cells due to a concern about the limits of miniaturization. The DNA double strands complementary hybridization rule is the cornerstone for DNA computing. Based on this, it uses well-designed DNA sequences with a variety of carefully selected parameters such as the position binding force of the double-strand formation to realize the chemical reaction of the DNA chain system for DNA computing.

The Origin:
Leonard Adleman of the university of southern California initially developed this field in 1994. Adleman put forward a DNA molecular biological calculation method based on the Hamilton graph and successfully achieved molecular computing in DNA solution for the first time. Adleman’s pioneering work opened a new field for computational science, which was of great significance and soon gained extensive attention from researchers in the field of mathematics, computer, biology, etc. In addition, other biological computing models, such as membrane computing, bacterial computing, evolutionary calculation, and virus calculation have been proposed and implemented. Adleman demonstrated a proof-of-concept use of DNA as a form of computation which solved the seven-point Hamilton path problem. In 1995, the idea for DNA-based memory was proposed by Eric Baum who conjectured that a vast amount of data can be stored in a tiny amount of DNA due to its ultra-high density. In Adleman's experiment, the Hamiltonian Path Problem was implemented notationally as “travelling salesman problem”. For this purpose, different DNA fragments were created, each one of them representing a city that had to be visited. Every one of these fragments is capable of a linkage with the other fragments created. These DNA fragments were produced and mixed in a test tube. Within seconds, the small fragments form bigger ones, representing the different travel routes. Through a chemical reaction, the DNA fragments representing the longer routes were eliminated. The remains are the solution to the problem, but overall, the experiment lasted a week.
Why is Molecular Computing Necessary?
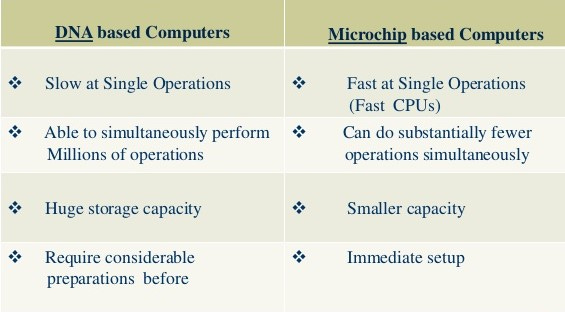
The power of the DNA computer lies in its massive parallel processing capability; when a chemical is added to the test tube it acts on every strand simultaneously. Because the average tube holds trillions of strands, that is a lot of computing going on at once. The core advantage of molecular computing is its potential to pack vastly more circuitry onto a microchip than silicon will ever be capable of—and to do it cheaply. DNA computing is a form of parallel computing in that it takes advantage of the many different molecules of DNA to try many different possibilities at once.
How it works
There are multiple methods for building a computing device based on DNA, each with its own advantages and disadvantages. Most of these build the basic logic gates (AND, OR, NOT) associated with digital logic from a DNA basis. Some of the different bases include DNAzymes, deoxyoligonucleotides, enzymes, toehold exchange. Here instead of using 0 and 1, we use high and low-intensity DNA signals as to when the DNA stand combines with its complementary strand(ie.A combines with the T in its complementary strand and G combines with the C in its complementary strand and vice-versa).
DNA computer is able to play tic-tac toe against a human player. The calculator consists of nine bins corresponding to the nine squares of the game. Each bin contains a substrate and various combinations of DNA enzymes. The substrate itself is composed of a DNA strand onto which was grafted a fluorescent chemical group at one end, and the other end, a repressor group. Fluorescence is only active if the molecules of the substrate are cut in half. The DNA enzymes simulate logical functions. For example, such a DNA will unfold if two specific types of DNA strand are introduced to reproduce the logic function AND.By default, the computer is considered to have played first in the central square. The human player starts with eight different types of DNA strands corresponding to the eight remaining boxes that may be played. To play box number i, the human player pours into all bins the strands corresponding to input #i. These strands bind to certain DNA enzymes present in the bins, resulting, in one of these bins, in the deformation of the DNA enzymes which binds to the substrate and cuts it. The corresponding bin becomes fluorescent, indicating which box is being played by the DNA computer. The DNA enzymes are divided among the bins in such a way as to ensure that the best the human player can achieve is a draw, as in real tic-tac-toe.
Pros and Cons:
The slow processing speed of a DNA computer (the response time is measured in minutes, hours or days, rather than milliseconds) is compensated by its potential to make a high amount of multiple parallel computations. This allows the system to take a similar amount of time for a complex calculation as for a simple one. This is achieved by the fact that millions or billions of molecules interact with each other simultaneously. However, it is much harder to analyze the answers given by a DNA computer than by a digital one.
An Article By: Biogen Team
DISCLAIMER
Shaastra TechShots’ publications contain information, opinions and data that Shaastra TechShots considers to be accurate based on the date of their creation and verified sources available at that time. It does not constitute either a personalized opinion or a general opinion of Shaastra or IIT Madras. The information provided comes from the best sources, however, Shaastra TechShots cannot be held responsible for any errors or omissions that may emerge.



